36 Hairpin Loops
36.1 Folding Free Energy Change
Hairpin loops of 4 or more nucleotides
The prediction of folding free energy changes for hairpins of 4 or more unpaired nucleotides is made with the following equation:
ΔG°37 hairpin (>3 nucleotides in loop) = ΔG°37 initiation (n) + ΔG°37 (terminal mismatch) + ΔG°37 (UU or GA first mismatch) + ΔG°37 (GG first mismatch) + ΔG°37 (special GU closure) + ΔG°37 penalty (all C loops)
In this equation, n is the number of nucleotides in loop, the terminal mismatch parameter is the sequence-dependent term for the first mismatch stacking on the terminal base pair, UU and GA first mismatches receive a bonus (not applied to AG first mismatches), GG first mismatches receive a bonus, the special GU closure term is applied only to hairpins in which a GU closing pair (not UG) is preceded by two Gs, and finally loops with all C nucleotides receive a penalty.
The penalty for all C loops of more than three unpaired nucleotides is predicted using a linear equation:
ΔG°37 penalty (all C loops; > 3 unpaired nucleotides) = An + B
Hairpin loops of 3 unpaired nucleotides
For hairpin loops of 3 nucleotides, the folding free energy change is estimated using:
ΔG°37 hairpin (3 unpaired nucleotides) = ΔG°37 initiation (3) + ΔG°37 penalty (all C loops)
As opposed to longer hairpin loops, hairpin loops of three nucleotides do not receive a sequence-dependent first mismatch term. All C hairpin loops of three nucleotides receive a stability penalty.
Special hairpin loops
There are hairpin loop sequences of 3, 4, and 6 nucleotides that have stabilities poorly fit by the model. These hairpins are assigned stabilities based on experimental data. Short hairpin loops
The nearest neighbor rules prohibit hairpin loops with fewer than 3 nucleotides.
36.2 Examples
6 nucleotide hairpin loop with no special stacking terms

ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(Hairpin Loop)
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(terminal mismatch) + ΔG°37 Hairpin initiation(6)
ΔG°37 = ΔG°37(CG followed by MU) + ΔG°37(MU followed by CG) + ΔG°37(CG followed by MU) + ΔG°37(MU followed by MM) + ΔG°37 Hairpin initiation(6)
ΔG°37 = -1.3 kcal/mol -1.9 kcal/mol -1.3 kcal/mol -0.8 kcal/mol + 5.4 kca/mol
ΔG°37 = 0.1 kcal/mol
Note that for unimolecular secondary structures, the helical intermolecular initiation does not appear.
5 nucleotide hairpin loop with a GG first mismatch

ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(Hairpin Loop)
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(terminal mismatch) + ΔG°37(GG first mismatch) + ΔG°37 Hairpin initiation(5)
ΔG°37 = ΔG°37(CG followed by AU) + ΔG°37(AU followed by CG) + ΔG°37(CG followed by AU) + ΔG°37 AU end penalty + ΔG°37(AU followed by GG) + ΔG°37(GG first mismatch) + ΔG°37 Hairpin initiation(5)
ΔG°37 = –2.1 kcal/mol – 2.2 kcal/mol – 2.1 kcal/mol + 0.5 kcal/mol – 0.8 kcal/mol – 0.8 kcal/mol + 5.7 kcal/mol
ΔG°37 = –1.9 kcal/mol
4 nucleotide special hairpin loop
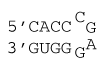
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(Hairpin Loop)
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(CcgagG)
ΔG°37 = ΔG°37(CG followed by AU) + ΔG°37(AU followed by CG) + ΔG°37(CG followed by CG) + ΔG°37(CcgagG)
ΔG°37 = –2.1 kcal/mol – 2.2 kcal/mol – 3.3 kcal/mol + 3.5 kcal/mol
ΔG°37 = -4.1 kcal/mol
6 nucleotide all C loop
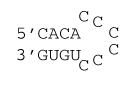
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(Hairpin Loop)
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(terminal mismatch) + ΔG°37 Hairpin initiation(6) + ΔG°37 penalty (all C loops)
ΔG°37 = ΔG°37(CG followed by AU) + ΔG°37(AU followed by CG) + ΔG°37(CG followed by AU) + ΔG°37 AU end penalty + ΔG°37(AU followed by CC) + ΔG°37 Hairpin initiation(6) + 6×A + B
ΔG°37 = –2.1 kcal/mol – 2.2 kcal/mol – 2.1 kcal/mol + 0.5 kcal/mol - 0.7 kcal/mol + 5.4 kcal/mol + 6×0.3 kcal/mol + 1.6 kcal/mol
ΔG°37 = 2.1 kcal/mol
5 nucleotide loop with special GU closure
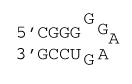
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(Hairpin Loop)
ΔG°37 = ΔG°37(Watson-Crick-Franklin Helix) + ΔG°37(terminal mismatch) + ΔG°37(GG first mismatch) + ΔG°37 Hairpin initiation(5) + ΔG°37 (special GU closure)
ΔG°37 = ΔG°37(CG followed by GC) + ΔG°37(GC followed by GC) + ΔG°37(GC followed by GU) + ΔG°37 GU end penalty + ΔG°37(GU followed by GG) + ΔG°37(GG first mismatch) + ΔG°37 Hairpin initiation(5) + ΔG°37 (special GU closure)
ΔG°37 = –2.36 kcal/mol – 3.26 kcal/mol – 1.53 kcal/mol + 0.45 kcal/mol – 0.8 kcal/mol – 0.8 kcal/mol + 5.7 kcal/mol – 2.2 kcal/mol
ΔG°37 = -4.8 kcal/mol
36.3 Parameter Tables
Length dependent initiation parameters are available in plain text for free energy changes. The plain text initiation tables include an extrapolation out to lengths of 30 nucleotides. Initiation parameters are based on experiments for sizes up to 9 nucleotides, but can be extrapolated to longer loops. For free energy changes, the extrapolation is ΔG°37 initiation (n>9) = ΔG°37 initiation (9) + 1.75 RT ln(n/9), where R is the gas constant and T is the absolute temperature. For enthalpy changes, ΔH°initiation (n>9) = ΔH°initiation (9).
The terminal mismatch tables are available in plain text for free energy change.
The table of special hairpin loops is available in plain text for free energy change for 3, 4, or 6 nucleotides. The special hairpin loop sequences include the identity of the closing basepair.
