9 Multibranch Loops
9.1 Folding Free Energy Change
Multibranch loops stabilities are predicted using the following equation:
ΔG°37 multibranch = ΔG°37 initiation + ΔG°37 stacking
where the stacking is the optimal configuration of dangling ends, terminal mismatches, or coaxial stacks, noting that a nucleotide or helix end can participate in only one of these favorable interactions.
Initiation is predicted using:
ΔG°37 initiation = a + b×[average asymmetry] + c×[number of branching helices] + ΔG°37 strain(three-way branching loops with fewer than two unpaired nucleotides)
where the average asymmetry is calculated as:
average asymmetry = min[2.0, mean difference in unpaired nucleotides on each side of each helix]
9.2 Folding Enthalpy Change
Similar to free energy change, multibranch loops enthalpy changes are predicted using the following equation:
ΔH°multibranch = ΔH°initiation + ΔH°stacking
where the stacking is the configuration of dangling ends, terminal mismatches, or coaxial stacks with lowest folding free energy change.
Initiation is predicted using an equation analagous to that folding free energy initiation:
ΔH°initiation = a + b×[average asymmetry] + c×[number of branching helices] + ΔH°strain(three-way branching loops with fewer than two unpaired nucleotides)
where the average asymmetry is calculated as:
average asymmetry = min[2.0, mean difference in unpaired nucleotides on each side of each helix]
9.3 Example
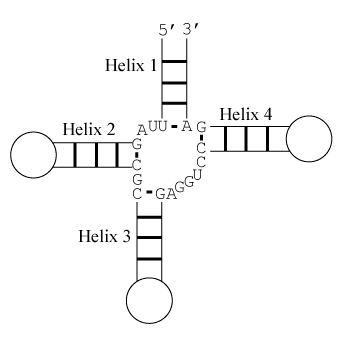
Free Energy Change
Prediction of Stacking
The predicted stacking configuration is the one with lowest free energy change. There are eight relevant configurations.
Configuration 1:
Helix 1 with \(3'\) dangling U, Helix 2 with terminal mismatch, Helix 3 with \(3'\) dangling A, and Helix 4 with \(5'\) dangling C
ΔG°37 = ΔG°37(UA with \(3'\) dangling U) + ΔG°37(CG followed by GA) + ΔG°37(GC with \(3'\) dangling A) + ΔG°37(GC with \(5'\) dangling C)
ΔG°37 = –0.1 kcal/mol – 1.4 kcal/mol – 1.1 kcal/mol – 0.3 kcal/mol
ΔG°37 = –2.9 kcal/mol
Configuration 2:
Helix 1 with \(3'\) dangling U, Helix 2 with \(5'\) dangling A, Helix 3 with terminal mismatch, and Helix 4 with \(5'\) dangling C
ΔG°37 = ΔG°37(UA with \(3'\) dangling U) + ΔG°37(CG with \(5'\) dangling A) + ΔG°37(GC followed by AG) + ΔG°37(GC with \(5'\) dangling C)
ΔG°37 = –0.1 kcal/mol – 0.2 kcal/mol – 1.3 kcal/mol – 0.3 kcal/mol
ΔG°37 = –1.9 kcal/mol
Configuration 3:
Helix 1 in flush coaxial stack with helix 4, Helix 2 with terminal mismatch, and Helix 3 with \(3'\) dangling A
ΔG°37 = ΔG°37(GC followed by AU) + ΔG°37(CG followed by GA) + ΔG°37(GC with \(3'\) dangling A)
ΔG°37 = –2.35 kcal/mol – 1.4 kcal/mol – 1.1 kcal/mol
ΔG°37 = –4.9 kcal/mol
Configuration 4:
Helix 1 in flush coaxial stack with helix 4, Helix 2 with \(5'\) dangling A, and Helix 3 with terminal mismatch
ΔG°37 = ΔG°37(GC followed by AU) + ΔG°37(CG with \(5'\) dangling A) + ΔG°37(GC followed by AG)
ΔG°37 = –2.35 kcal/mol – 0.2 kcal/mol – 1.3 kcal/mol
ΔG°37 = –3.9 kcal/mol
Configuration 5:
Helix 1 with \(3'\) dangling U, Helix 2 in mismatch–mediated coaxial stack with helix 3 with GA intervening mismatch, and Helix 4 with \(5'\) dangling C
ΔG°37 = ΔG°37(UA with \(3'\) dangling U) + ΔG°37(CG followed by GA) + ΔG°37(Discontinuous Backbone Stack) + ΔG°37(GC with \(5'\) dangling C)
ΔG°37 = –0.1 kcal/mol – 1.4 kcal/mol – 2.1 kcal/mol – 0.3 kcal/mol
ΔG°37 = –3.9 kcal/mol
Configuration 6:
Helix 1 with \(3'\) dangling U, Helix 2 in mismatch–mediated coaxial stack with helix 3 with AG intervening mismatch, and Helix 4 with \(5'\) dangling C
ΔG°37 = ΔG°37(UA with \(3'\) dangling U) + ΔG°37(Discontinuous Backbone Stack) + ΔG°37(GC followed by AG) + ΔG°37(GC with \(5'\) dangling C)
ΔG°37 = –0.1 kcal/mol – 2.1 kcal/mol – 1.3 kcal/mol – 0.3 kcal/mol
ΔG°37 = –3.8 kcal/mol
Configuration 7:
Helix 1 in flush coaxial stack with helix 4 and Helix 2 in mismatch–mediated coaxial stack with helix 3 with GA intervening mismatch
ΔG°37 = ΔG°37(GC followed by AU) + ΔG°37(CG followed by GA) + ΔG°37(Discontinuous Backbone Stack)
ΔG°37 = –2.35 kcal/mol – 1.4 kcal/mol – 2.1 kcal/mol
ΔG°37 = –5.9 kcal/mol
Configuration 8:
Helix 1 in flush coaxial stack with helix 4 and Helix 2 in mismatch–mediated coaxial stack with helix 3 with AG intervening mismatch
ΔG°37 = ΔG°37(GC followed by AU) + ΔG°37(Discontinuous Backbone Stack) + ΔG°37(GC followed by AG)
ΔG°37 = –2.35 kcal/mol – 2.1 kcal/mol – 1.3 kcal/mol
ΔG°37 = –5.8 kcal/mol
Configuration 7 has the lowest folding free energy change of –5.9 kcal/mol.
Initiation Free Energy Change
ΔG°37 initiation = a + b×[average asymmetry] + c×[number of branching helices] + ΔG°37 strain(three–way branching loops with fewer than two unpaired nucleotides)
ΔG°37 initiation = 9.25 kcal/mol + (0.91 kcal/mol)×[average asymmetry] + (–0.63 kcal/mol)×[4]
Average asymmetry = min[2.0,(2+1+4+5)/4] = min[2.0,3.0] = 2.0
ΔG°37 initiation = 9.25 kcal/mol + (0.91 kcal/mol)×[2] + (–0.63 kcal/mol)×[4]
ΔG°37 initiation = 8.6 kcal/mol
Total Folding Free Energy Change
ΔG°37 multibranch loop = ΔG°37 initiation + ΔG°37 stacking = 8.6 kcal/mol – 5.9 kcal/mol = 2.7 kcal/mol
9.3.1 Enthalpy Change
Prediction of Stacking
The stacking configuration is fixed by the prediction of folding free energy change and is configuration 7 above.
ΔH°= ΔH°(GC followed by AU) + ΔH°(CG followed by GA) + ΔH°(Discontinuous Backbone Stack)
ΔH° = –12.44 kcal/mol – 8.2 kcal/mol – 8.46 kcal/mol
ΔH° = –29.1 kcal/mol
Initiation Enthalpy Change
ΔH°initiation = a + b×[average asymmetry] + c×[number of branching helices] + ΔH°strain(three–way branching loops with fewer than two unpaired nucleotides)
ΔH°initiation = 38.9 kcal/mol + (12.9 kcal/mol)×[average asymmetry] + (–11.9 kcal/mol)×[4]
Average asymmetry = min[2.0,(3+2+4+5)/4] = min[2.0,3.0] = 2.0
ΔH°initiation = 38.9 kcal/mol + (12.9 kcal/mol)×[2] + (–11.9 kcal/mol)×[4]
ΔH°initiation = 17.1 kcal/mol
Total Folding Enthalpy Energy Change
ΔH°multibranch loop = ΔH°initiation + ΔH°stacking = 17.1 kcal/mol – 29.1 kcal/mol = –12.0 kcal/mol
Note that helices 1 and 2 are separated by two unpaired nucleotides and cannot stack coaxially. Similarly, helices 3 and 4 are too distant to stack coaxially. Also note that coaxial stacking is only allowed between adjacent helices and hence, for example, helices 1 and 3 cannot stack coaxially.
9.4 Parameter Tables
Tables of parameters are available in html format.
9.5 References
The multibranch loop nearest neighbor parameters for initiation free energy change were reported in:
Mathews, D.H. and Turner, D.H. (2002) Experimentally derived nearest neighbor parameters for the stability of RNA three- and four-way multibranch loops. Biochemistry, 41, 869-880.
The enthalpy change parameters were reported in:
Lu, Z.J., Turner, D.H. and Mathews, D.H. (2006) A set of nearest neighbor parameters for predicting the enthalpy change of RNA secondary structure formation. Nucleic Acids Res., 34 4912 - 4924.
The experimental data for the fit of the parameters were taken from:
- Diamond, J.M., Turner, D.H. and Mathews, D.H. (2001) Thermodynamics of three-way multibranch loops in RNA. Biochemistry, 40, 6971-6981.
- Mathews, D.H. and Turner, D.H. (2002) Experimentally derived nearest neighbor parameters for the stability of RNA three- and four-way multibranch loops. Biochemistry, 41, 869-880.
